Soil microbiomes and their arsenic functional genes in chronically high-arsenic contaminated soils
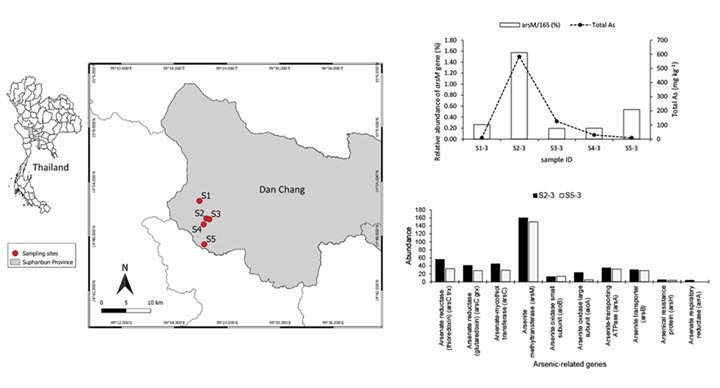
Microbial arsenic transformations play essential roles in controlling pollution and ameliorating risk. This study combined high-throughput sequencing and PCR-based approaches targeting both the 16 S rRNA and arsenic functional genes to investigate the temporal and spatial dynamics of the soil microbiomes impacted by high arsenic contamination (9.13 to 911.88 mg/kg) and to investigate the diversity and abundance of arsenic functional genes in soils influenced by an arsenic gradient. The results showed that the soil microbiomes were relatively consistent and mainly composed of Actinobacteria (uncultured Gaiellales and an unknown_67 − 14 bacterium), Proteobacteria, Firmicutes (particularly, Bacillus), Chloroflexi, and Acidobacteria (unknown_Subgroup_6). Although a range of arsenic functional genes (e.g., arsM, arsC, arrA, and
aioA) were identified by shotgun metagenomics, only the arsM gene was detected by the PCR-based method. The relative abundance of the arsM gene accounted for 0.20%–1.57% of the total microbial abundance. Combining all analyses, arsenic methylation mediated by the arsM gene was proposed to be a key process involved in the arsenic biogeochemical cycle and
mitigation of arsenic toxicity. This study advances our knowledge about arsenic mechanisms over the long-term in highly contaminated soils.
สนับสนุน SDGs เป้าหมาย 15 ปกป้อง ฟื้นฟู และสนับสนุนการใช้ระบบนิเวศบนบกอย่างยั่งยืน จัดการป่าไม้อย่างยั่งยืน ต่อสู้การกลายสภาพเป็นทะเลทราย หยุดการเสื่อมโทรมของที่ดินและฟื้นสภาพกลับมาใหม่ และหยุดการสูญเสียความหลากหลายทางชีวภาพ
อ้างอิง
Sonthiphand, P., Rueangmongkolrat, N., Uthaipaisanwong, P. et al. Soil Microbiomes and their Arsenic Functional Genes in Chronically High-Arsenic Contaminated Soils. Bull Environ Contam Toxicol 112, 49 (2024). https://doi.org/10.1007/s00128-024-03866-1